SoDCoD
a comprehensive database of Cu/Zn Superoxide Dismutase Conformational Diversity caused by ALS-linked gene mutations and other perturbations
Cu/Zn Superoxide Dismutase (SOD1) is one of the causative genes for amyotrophic lateral sclerosis (ALS), and over 180 distinct mutations accompanied by amino acid changes have been reported in ALS patients. We present SoDCoD, a database collecting our comprehensive biochemical analyses for the structural changes of SOD1 caused by ALS-linked gene mutations and other perturbations.
We previously carried out a comprehensive analysis of all the SOD1 mutants on the following two points:
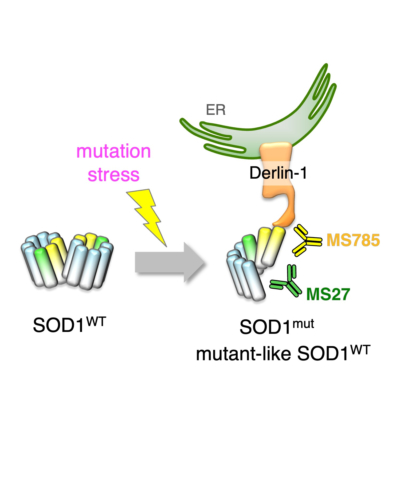
- Interaction with Derlin-1(CT4) We reported that most SOD1 mutants bind to the carboxyl-terminal region of Derlin-1 [termed Derlin-1(CT4)], a component of the endoplasmic reticulum-associated degradation (ERAD) machinery, and this interaction triggers ER stress-dependent motor neuron death.
- Conformatinal alterlation We reported that most SOD1 mutants have a constitutively exposed Derlin-1-binding region (DBR) that is occluded in the wild-type protein. We developed monoclonal antibodies, MS785 and MS27, which recognizes DBR-exposed SOD1. A systematic immunoprecitipation analysis revealed that most SOD1 mutants expose the DBR.
In addition, we found that zinc deficiency induces a conformational alteration in wild-type SOD1. We performed an siRNA screening to identify genes that were involved in the proteostasis of conformationally altered wild-type SOD1. We catalogued candidate genes in the screen on this database.
This database provides indispensable information for significant progress in medical research, diagnosis, and treatment of ALS targeting conformational alteration in SOD1.
This version 1.0 contains information on the properties of 188 types of SOD1 mutants, including structural changes and binding to Derlin-1, as well as a set of genes that contribute to the proteostasis of mutant-like wild-type SOD1.
How to Use
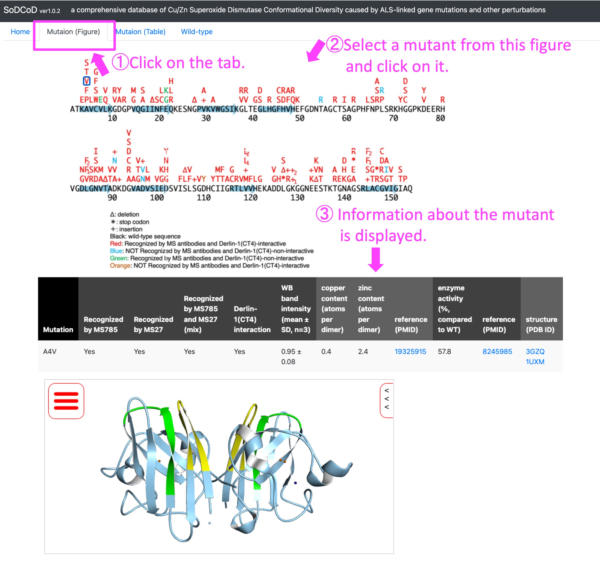
- Obtain information on the structural properties of SOD1 mutants from a schematic diagram of SOD1 mutations
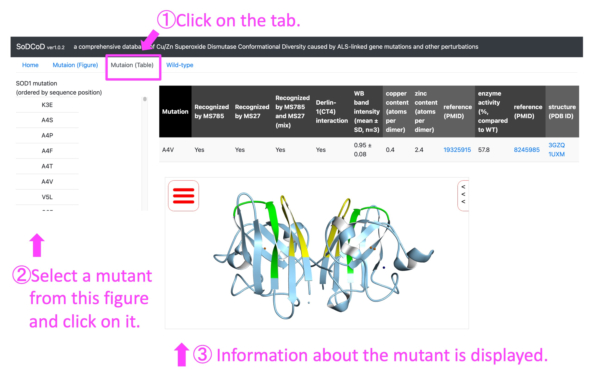
- Obtain information on the structural properties of SOD1 mutants from a list of mutations
- Obtain information on genes involved in the proteostasis of mutant-like wild-type SOD1

All raw data are deposited on GitHub. GitHub Repository
References
-
Fujisawa T, Homma K, Yamaguchi N, Kadowaki H, Tsuburaya N, Naguro I, Matsuzawa A, Takeda K, Takahashi Y, Goto J, Tsuji S, Nishitoh H, Ichijo H
A novel monoclonal antibody reveals a conformational alteration shared by amyotrophic lateral sclerosis-linked SOD1 mutants
Ann. Neurol., 72, 739-749 (2012)
-
Fujisawa T, Yamaguchi N, Kadowaki H, Tsukamoto Y, Tsuburaya N, Tsubota A, Takahashi H, Naguro I, Takahashi Y, Goto J, Tsuji S, Nishitoh H, Homma K, Ichijo H
A systematic immunoprecipitation approach reinforces the concept of common conformational alterations in amyotrophic lateral sclerosis-linked SOD1 mutants
Neurobiol. Dis., 82, 478-486 (2015)
-
Homma K, Takahashi H, Tsuburaya N, Naguro I, Fujisawa T and Ichijo H
Genome-wide siRNA screening reveals that DCAF4-mediated ubiquitination of optineurin stimulates autophagic degradation of Cu/Zn superoxide dismutase
J. Biol. Chem., 295, 3148-3158 (2020)
Contributors
Takao Fujisawa -Website development-
Yurika Momozawa -Website development-
Hideki Nishitoh -Graphic design on the left side in the home tab-
Questions and Comments
Please contact Takao Fujisawa (fujisawa@mol.f.u-tokyo.ac.jp)
Sequence Position Order
Browse ALS-linked SOD1 variants in sequence order. Select any mutation to update the detail table and structure view.
| Mutation | Recognized by MS785 | Recognized by MS27 | Recognized by MS785 and MS27 (mix) | Derlin-1(CT4) interaction | WB band intensity (mean ± SD, n=3) | copper content (atoms per dimer) | zinc content (atoms per dimer) | reference (PMID) | enzyme activity (%, compared to WT) | reference (PMID) | structure (PDB ID) |
|---|
| Mutation | Recognized by MS785 | Recognized by MS27 | Recognized by MS785 and MS27 (mix) | Derlin-1(CT4) interaction | WB band intensity (mean ± SD, n=3) | copper content (atoms per dimer) | zinc content (atoms per dimer) | reference (PMID) | enzyme activity (%, compared to WT) | reference (PMID) | structure (PDB ID) |
|---|